Methods of calculating protein hydrophobicity and their application in developing correlations to predict hydrophobic interaction chromatography retention - ScienceDirect

Characterizing hydrophobicity of amino acid side chains in a protein environment via measuring contact angle of a water nanodroplet on planar peptide network | PNAS

Clustering of Large Hydrophobes in the Hydrophobic Core of Two-stranded α-Helical Coiled-Coils Controls Protein Folding and Stability* - Journal of Biological Chemistry
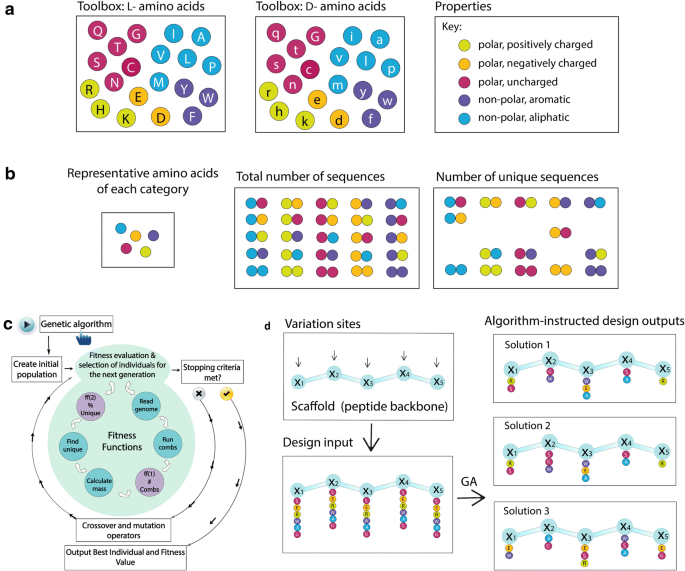
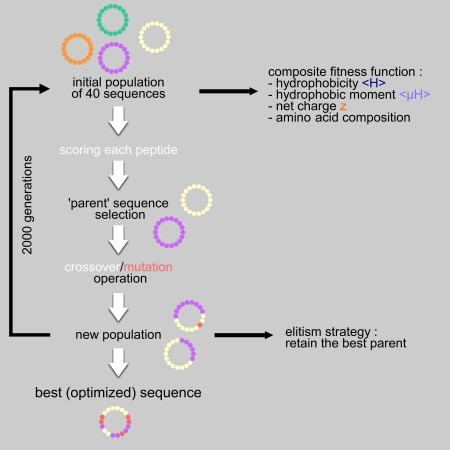
Algorithm-supported, mass and sequence diversity-oriented random peptide library design | Journal of Cheminformatics | Full Text

What can machine learning do for antimicrobial peptides, and what can antimicrobial peptides do for machine learning? | Interface Focus

Equation 1 was used to determine effective hydrophobicity. The helical... | Download Scientific Diagram

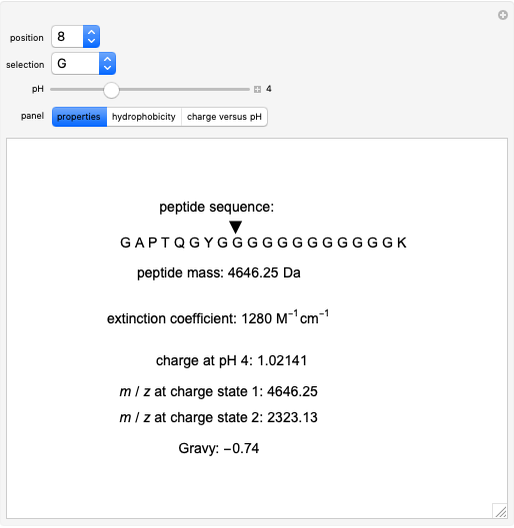
Bachem Group - Calculate your Peptide. Our Peptide Calculator is an online tool that calculates molecular weight, net charge, isoelectric point, and average hydrophobicity for your peptide sequence. Try it out. http://www.bachem.com/service-support ...

Characterizing hydrophobicity of amino acid side chains in a protein environment via measuring contact angle of a water nanodroplet on planar peptide network | PNAS
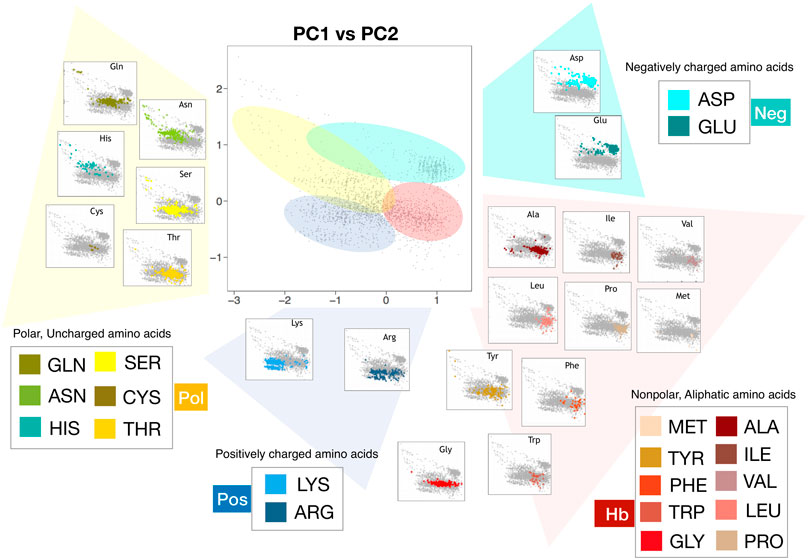
Frontiers | Characterizing Hydropathy of Amino Acid Side Chain in a Protein Environment by Investigating the Structural Changes of Water Molecules Network

Mean hydrophobicity, hydrophobic moment, net charge and helical wheel... | Download Scientific Diagram

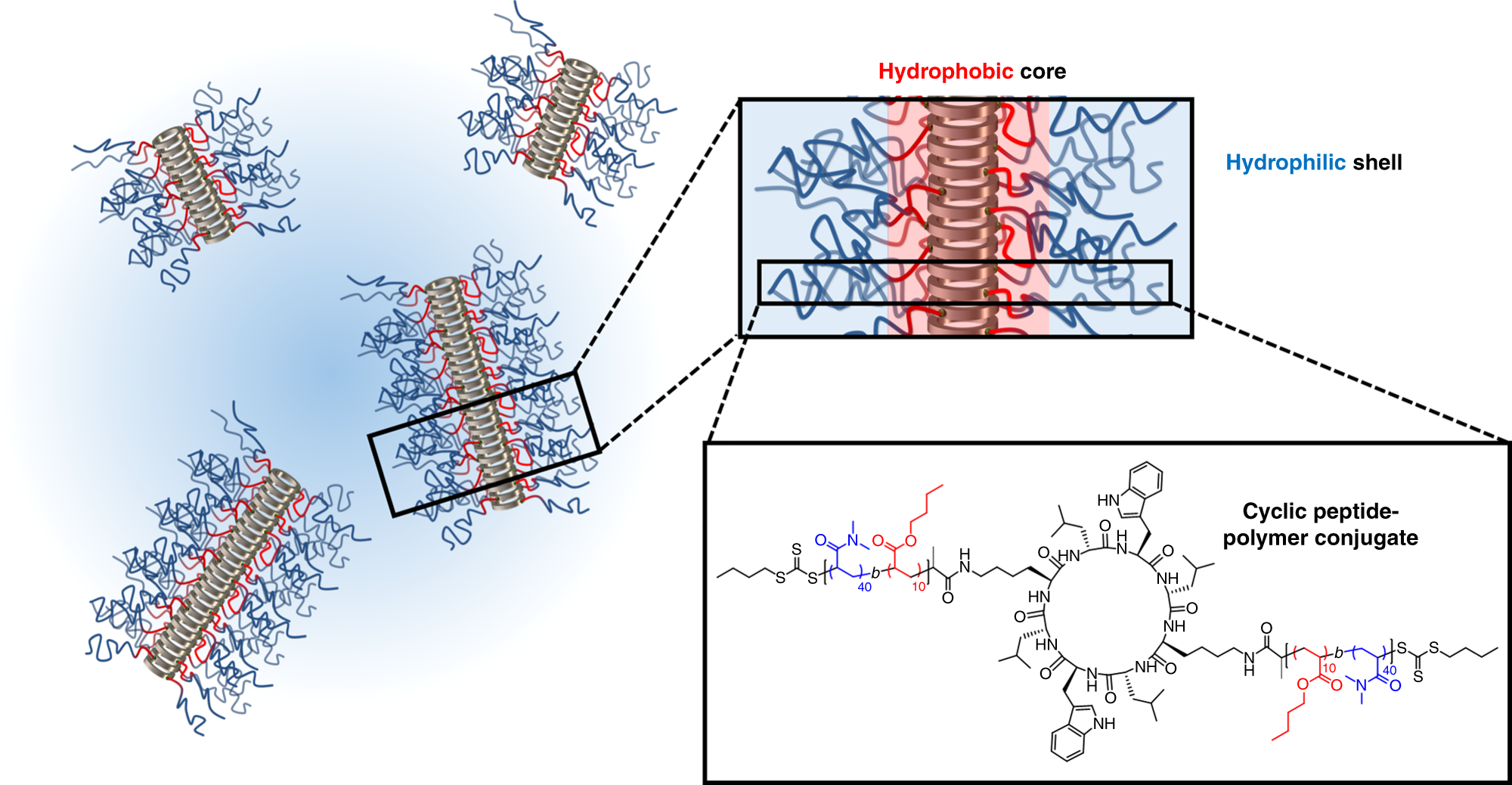
Peptide-polysaccharide conjugates with adjustable hydrophilicity/ hydrophobicity as green and pH sensitive emulsifiers - ScienceDirect

Efficient transdermal delivery of functional protein cargoes by a hydrophobic peptide MTD 1067 | Scientific Reports

Binding of cationic peptides (KX)4K to DPPG bilayers. Increasing the hydrophobicity of the uncharged amino acid X drives formation of membrane bound β-sheets: A DSC and FT-IR study - ScienceDirect

Space-filling model of parent peptide V13K L . Hydrophobic amino acids... | Download Scientific Diagram

Analysis of hydrophobic and hydrophilic moments of short penetrating peptides for enhancing mitochondrial localization: prediction and validation - Pirisinu - 2019 - The FASEB Journal - Wiley Online Library
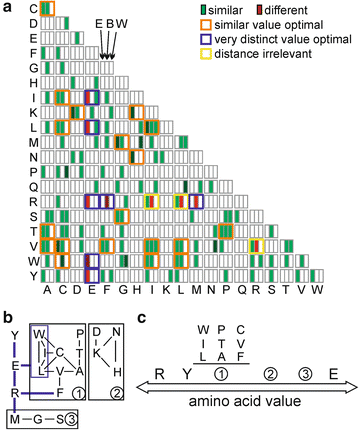
50 years of amino acid hydrophobicity scales: revisiting the capacity for peptide classification | Biological Research | Full Text

Solubility test of NSP and NSP r. (A) Peptide models computed by the... | Download Scientific Diagram

Peptide Retention Standards and Hydrophobicity Indexes in Reversed-Phase High-Performance Liquid Chromatography of Peptides | Analytical Chemistry